Chapter 4 Tables, conditionals and loops
Last updated: 2020-08-12 00:35:51
Aims
Our aims in this chapter are:
- Learn to work with
data.frame, the data structure used to represent tables in R - Learn several automation methods for controlling code execution and automation in R:
- Conditionals
- Loops
- The
applyfunction
- Join between tables
4.1 Tables
4.1.1 What is a data.frame?
A table in R is represented using the data.frame class. A data.frame is basically a collection of vectors comprising columns, all of the same length but possibly of different types. Conventionally:
- Each row represents an observation, with values possibly of a different type for each variable
- Each column represents a variable, with values of the same type
For example, the file rainfall.csv, which we are going to work with later on (Section 4.4), contains a table with information about meteorological stations in Israel (Figure 4.1). The table rows correspond to 169 meteorological stations, while table columns refer to different variables: station name (character), station coordinates (numeric), average monthly rainfall amounts (numeric), etc.
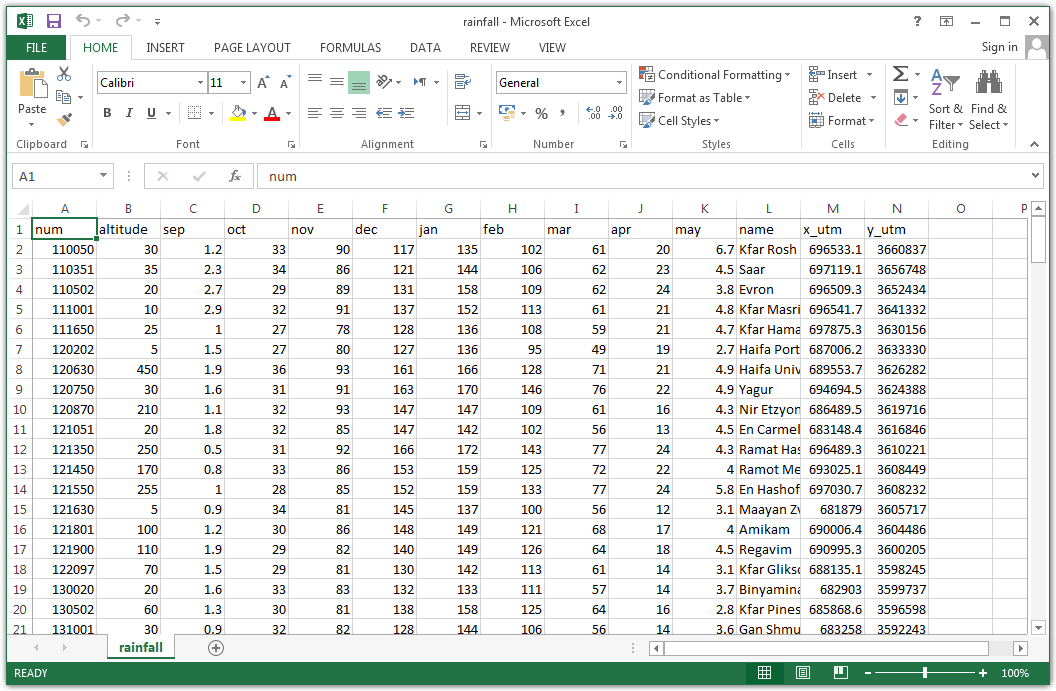
Figure 4.1: rainfall.csv opened in Excel
4.1.2 Creating a data.frame
A data.frame can be created with the data.frame function, given one or more vectors which become columns. The stringAsFactors=FALSE argument prevents the conversion of text columns to factor, which is what we usually want14.
For example, the following expression creates a table with four properties for three railway stations in Israel. The properties are:
name—Station namecity—The city where the station is locatedlines—The number of railway lines that go through the stationpiano—Does the station have a piano?
dat = data.frame(
name = c("Beer-Sheva Center", "Beer-Sheva University", "Dimona"),
city = c("Beer-Sheva", "Beer-Sheva", "Dimona"),
lines = c(4, 5, 1),
piano = c(FALSE, TRUE, FALSE),
stringsAsFactors = FALSE
)
dat
## name city lines piano
## 1 Beer-Sheva Center Beer-Sheva 4 FALSE
## 2 Beer-Sheva University Beer-Sheva 5 TRUE
## 3 Dimona Dimona 1 FALSE4.1.3 Interactive view of a data.frame
The View function opens an interactive view of a data.frame. When using RStudio, the view also has sort and filter buttons (Figure 4.2). Note that sorting or filtering the view have no effect on the object.
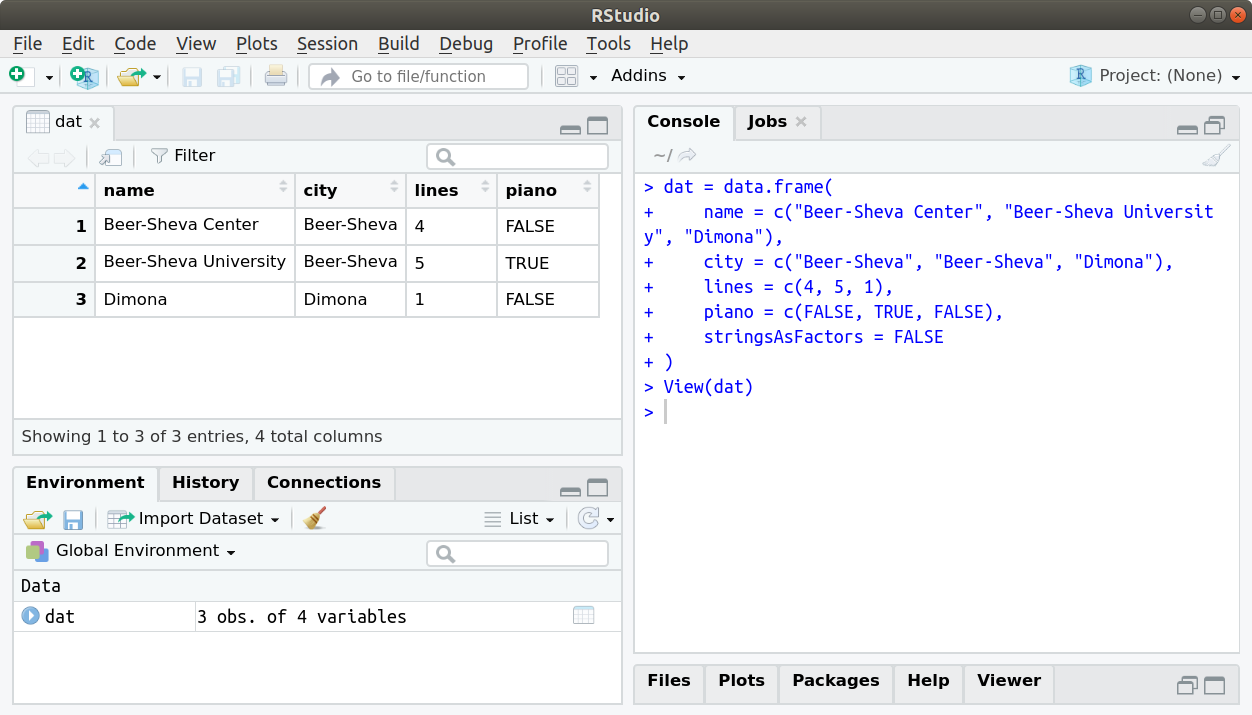
Figure 4.2: Table view in Rstudio
4.1.4 data.frame properties
4.1.4.1 Dimensions
Unlike a vector, which is one-dimensional—the number of elements is obtained with length (Section 2.3.4)—a data.frame is two-dimensional. We can get the number of rows and number of columns in a data.frame with nrow and ncol, respectively:
As an alternative, we can get both the number of rows and columns (in that order!), as a vector of length 2, with dim:
4.1.4.2 Row and column names
Any data.frame object also has row and column names, which we can get with rownames and colnames, respectively. Row names are usually meaningless, e.g., composed of consecutive numbers by default:
Conversely, column names are usually meaningful variable names:
We can also set row or column names by assigning new values to these properties, or to subsets thereof. For example, here is how we can change the first column name:
colnames(dat)[1] = "STATION_NAME"
dat
## STATION_NAME city lines piano
## 1 Beer-Sheva Center Beer-Sheva 4 FALSE
## 2 Beer-Sheva University Beer-Sheva 5 TRUE
## 3 Dimona Dimona 1 FALSEand revert to the previous name:
colnames(dat)[1] = "name"
dat
## name city lines piano
## 1 Beer-Sheva Center Beer-Sheva 4 FALSE
## 2 Beer-Sheva University Beer-Sheva 5 TRUE
## 3 Dimona Dimona 1 FALSEThe str function gives a summary of any object structure. For data.frame objects, str lists the dimensions, as well as column names, types and (first few) values in each column:
4.1.5 data.frame subsetting
4.1.5.1 Introduction
A data.frame subset can be obtained with the [ operator, which we are familiar with from vector subsetting (Sections 2.3.3, 2.3.8). A data.frame is a two-dimensional object, therefore the index is composed of two vectors:
- The first vector refers to rows
- The second vector refers to columns
Each of these vectors can be one of the following types:
numeric—Specifying the indices of rows/columns to retaincharacter—Specifying the names of rows/columns to retainlogical—Specifying whether to retain each row/column
Either the rows or the column index can be omitted, in which case we get all rows or all columns, respectively.
4.1.5.2 Numeric index
Here are several examples of subsetting with a numeric index:
4.1.5.3 The drop parameter
The subset operator [ accepts an additional logical argument drop. The drop argument determines whether we would like to simplify the resulting subset to a simpler data structure, when possible, “dropping” the more complex class (drop=TRUE, the default), or whether we would like to always keep the subset in its original class (drop=FALSE).
For example, a subset that conatains a single data.frame column can be returned as:
- A vector (
drop=TRUE, the default) - A
data.frame(drop=FALSE)
For example, a subset that contains a single column is, by default, simplified to a vector:
unless we specify drop=FALSE, in which case it remains a data.frame:
Why do you think simplification works when taking a subset with a single column, but not on a subset with a single row?
4.1.5.4 Character index
We can also use a character index to specify the names of rows and/or columns to retain in the subset:
dat[, c("name", "city")]
## name city
## 1 Beer-Sheva Center Beer-Sheva
## 2 Beer-Sheva University Beer-Sheva
## 3 Dimona DimonaThe $ operator is a shortcut for getting a single column, by name, from a data.frame:
4.1.5.5 Logical index
The third option for a data.frame index is a logical vector, specifying whether to retain each row or column. Most commonly it is used to filter data.frame rows, based on the values of one or more columns. For example:
dat[dat$city == "Beer-Sheva", ]
## name city lines piano
## 1 Beer-Sheva Center Beer-Sheva 4 FALSE
## 2 Beer-Sheva University Beer-Sheva 5 TRUEdat[dat$city == "Beer-Sheva" & !dat$piano, ]
## name city lines piano
## 1 Beer-Sheva Center Beer-Sheva 4 FALSELet’s go back to the Kinneret example from Chapter 3:
may = c(
-211.92,-208.80,-208.84,-209.12,-209.01,-209.60,-210.24,-210.46,-211.76,
-211.92,-213.13,-213.18,-209.74,-208.92,-209.73,-210.68,-211.10,-212.18,
-213.26,-212.65,-212.37
)
nov = c(
-212.79,-209.52,-209.72,-210.94,-210.85,-211.40,-212.01,-212.25,-213.00,
-213.71,-214.78,-214.34,-210.93,-210.69,-211.64,-212.03,-212.60,-214.23,
-214.33,-213.89,-213.68
)
year = 1991:2011We already know how to combine the vectors into a data.frame:
kineret = data.frame(year, may, nov)
kineret
## year may nov
## 1 1991 -211.92 -212.79
## 2 1992 -208.80 -209.52
## 3 1993 -208.84 -209.72
## 4 1994 -209.12 -210.94
## 5 1995 -209.01 -210.85
## 6 1996 -209.60 -211.40
## 7 1997 -210.24 -212.01
## 8 1998 -210.46 -212.25
## 9 1999 -211.76 -213.00
## 10 2000 -211.92 -213.71
## 11 2001 -213.13 -214.78
## 12 2002 -213.18 -214.34
## 13 2003 -209.74 -210.93
## 14 2004 -208.92 -210.69
## 15 2005 -209.73 -211.64
## 16 2006 -210.68 -212.03
## 17 2007 -211.10 -212.60
## 18 2008 -212.18 -214.23
## 19 2009 -213.26 -214.33
## 20 2010 -212.65 -213.89
## 21 2011 -212.37 -213.68Using a logical index we can get a subset of years when the Kinneret level in November was less than -213. The following expression is identical to the one we used when working with separate vectors (Section 3.1.3), except for the dat$ part, which specifies that we refer to data.frame columns:
When operating on a data.frame, we can also get a subset with data from all columns for those selected years, as follows:
kineret[kineret$nov < -213, ]
## year may nov
## 10 2000 -211.92 -213.71
## 11 2001 -213.13 -214.78
## 12 2002 -213.18 -214.34
## 18 2008 -212.18 -214.23
## 19 2009 -213.26 -214.33
## 20 2010 -212.65 -213.89
## 21 2011 -212.37 -213.68What are the differences between the last two expressions? What is the reason for those differences?
4.1.6 Creating new columns
Assignment into a column which does not exist adds a new column. For example, here is how we can add a new column named d_nov, containing consecutive differences between values in the nov column (Section 3.2.3):
kineret$d_nov = c(NA, diff(kineret$nov))
kineret
## year may nov d_nov
## 1 1991 -211.92 -212.79 NA
## 2 1992 -208.80 -209.52 3.27
## 3 1993 -208.84 -209.72 -0.20
## 4 1994 -209.12 -210.94 -1.22
## 5 1995 -209.01 -210.85 0.09
## 6 1996 -209.60 -211.40 -0.55
## 7 1997 -210.24 -212.01 -0.61
## 8 1998 -210.46 -212.25 -0.24
## 9 1999 -211.76 -213.00 -0.75
## 10 2000 -211.92 -213.71 -0.71
## 11 2001 -213.13 -214.78 -1.07
## 12 2002 -213.18 -214.34 0.44
## 13 2003 -209.74 -210.93 3.41
## 14 2004 -208.92 -210.69 0.24
## 15 2005 -209.73 -211.64 -0.95
## 16 2006 -210.68 -212.03 -0.39
## 17 2007 -211.10 -212.60 -0.57
## 18 2008 -212.18 -214.23 -1.63
## 19 2009 -213.26 -214.33 -0.10
## 20 2010 -212.65 -213.89 0.44
## 21 2011 -212.37 -213.68 0.214.2 Flow control
4.2.1 Introduction
The default execution mode is to let the computer execute all expressions in the same order they are given in the code. Flow control commands are a way to modify the sequence of code execution. We will learn two flow control operators, from two flow control categories:
ifandelse—A conditional, conditioning the execution of codefor—A loop, executing code more than once
4.2.2 Conditionals
The purpose of the conditional is to condition the execution of code. An if-else conditional in R contains the following components:
- The
ifkeyword - A condition
- Code to be executed if the condition is
TRUE - The
elsekeyword (optional) - Code to be executed if the condition is
FALSE(optional)
The condition needs to be evaluated to a logical vector of length 1, containing either TRUE or FALSE. If the condition is TRUE, then the code section after if is executed. If the condition is FALSE, then the code section after else (when present) is executed.
Here is the syntax of a conditional with if:
and here is the syntax of a conditional with if and the (optional) else:
The following examples demonstrate how the expression after if is executed when the condition is TRUE:
When the condition is FALSE—nothing happens:
Now let’s also add a second expression after else. The first code section is still executed when the condition is TRUE:
When the condition is FALSE, however, the second code section is executed:
Conditionals are frequently used when our code branches into two scenarios, depending on the value of a particular object. For example, we can use a conditional to define (Section 3.3) our own version of the abs function (Section 2.3.4):
Let’s check if our custom function abs2 works as expected:
Seems like it does, at least for arguments that are vectors of length 1.
What happens when the argument of
abs2is of length >1? What is the reason for the warning we get and why does the function return a “wrong” answer when the first element is negative?
4.2.3 Loops
A loop is used to execute a given code section more than once. The number of times the code is executed is determined in different ways in different types of loops. In a for loop, the number of times the code is executed is determined in advance, based on the length of a vector passed to the loop. The code is executed once for each element of the vector. In each “round”, the current element is assigned to a variable which we can use in the loop code.
A for loop is composed of the following parts:
- The
forkeyword - The variable name
symbolgetting the current vector value - The
inkeyword - The vector
sequence - A code section
expressions
Here is the syntax of a for loop:
Note that the constant keywords are just for and in. All other components (symbol, sequence and expressions) are varying, and it is up to us to choose their values.
Here is an example of a for loop:
What has happened? The expression print(i) was executed 5 times, according to the length of the vector 1:5. Each time, i got the next value of 1:5 and the code section printed that value on screen.
The vector defining the for loop does not necessarily need to be numeric. For example:
Here, expression print(b) was executed 3 times, according to the length of the vector c("Test", "One", "Two"). Each time, b got the next value of the vector and the code section printed b on screen.
In case the vector is numeric, it does not necessarily need to be composed of consecutive:
Again, the expression print(i) was executed 3 times, now according to the length of the vector c(1,15,3). Each time, i got the next value of the vector and the code section printed i on screen.
The code section even does not have to use the current value of the vector:
Here, the expression print("A") was executed 5 times, according to the length of the vector 1:5. Each time, the code section printed the fixed value "A" on screen.
The following for loop prints each of the numbers from 1 to 10 multiplied by 5:
for(i in 1:10) print(i * 5)
## [1] 5
## [1] 10
## [1] 15
## [1] 20
## [1] 25
## [1] 30
## [1] 35
## [1] 40
## [1] 45
## [1] 50How can we print a multiplication table for 1-10, using a
forloop, as shown below?
## [1] 1 2 3 4 5 6 7 8 9 10
## [1] 2 4 6 8 10 12 14 16 18 20
## [1] 3 6 9 12 15 18 21 24 27 30
## [1] 4 8 12 16 20 24 28 32 36 40
## [1] 5 10 15 20 25 30 35 40 45 50
## [1] 6 12 18 24 30 36 42 48 54 60
## [1] 7 14 21 28 35 42 49 56 63 70
## [1] 8 16 24 32 40 48 56 64 72 80
## [1] 9 18 27 36 45 54 63 72 81 90
## [1] 10 20 30 40 50 60 70 80 90 100As another example of using a for loop, we can write a function named x_in_y. The function accepts two vectors x and y. For each element in x the function checks whether it is found in y. It returns a logical vector of the same length as x.
Here is an example of how the function is supposed to work:
In plain terms, what we need to do is to go over the elements of x, each time checking whether the current element is equal to any of the elements in y. This is exactly the type of operation where a for loop comes in handy. We can use a for loop to check if each element in x is contained in y, as follows:
Inside a function, rather then printing we would like to “collect” the results into a vector. There are at least two ways to do it. One way it to start from NULL, which specifies an empty object (Section 1.3.5), then consecutively add new elements with c:
Another way is to start from a vector composed of NA with the right length (rep(NA, length(x)))), then fill-in the results using assignment:
x_in_y = function(x, y) {
result = rep(NA, length(x))
for(i in 1:length(x)) result[i] = any(x[i] == y)
result
}Situations when we need to go over subsets of a dataset, process those subsets, then combine the results back to a single object, are very common in data processing. A for loop is the default approach for such tasks, unless there is a “shortcut” that we may prefer, such as the apply function (Section 4.5). For example, we will come back to for loops when separately processing raster layers for several time periods (Section 11.3.2).
4.3 The %in% operator
In fact, we don’t need to write a function such as x_in_y (Section 4.2.3) ourselves; we can use the %in% operator. The %in% operator, with an expression x %in% y, returns a logical vector indicating the presence of each element of x in y. For example:
4.4 Reading tables from a file
4.4.1 Using read.csv
In addition to creating a table with data.frame (Section 4.1.2), we can read an existing table from disk (Figure 4.3), such as from a Comma-Separated Values (CSV) file.
Figure 4.3: Reading a file takes information from the mass storage and loads it into the RAM
In the next few examples we will work with the CSV file named rainfall.csv (Figure 4.1). This file contains a table with average monthly (September through May) rainfall data, based on the period of 1980-2010, in 169 meteorological stations in Israel. The table also contains station name, station number, elevation and X-Y coordinates.
We can read a CSV file using the read.csv function, given the file path. For example, here is how we can read the CSV file rainfall.csv assuming it is located in the C:\Data2 directory:
Note that the separating character is / or \\, not the familiar \, so either of the above expressions can be used to read the file. In case the file path uses the incorrect separator \, we get an error:
read.csv("C:\Data2\rainfall.csv")
## Error: '\D' is an unrecognized escape in character string starting ""C:\D"In case the file does not exist we get a different error:
read.csv("C:\\Data2\\rainfall.csv")
## Warning in file(file, "rt"): cannot open file 'C:\Data2\rainfall.csv': No such
## file or directory
## Error in file(file, "rt"): cannot open the connectionThe stringsAsFactors parameter of read.csv—which we already met in Section 4.1.2—determines whether text columns are converted to factor (the default is TRUE). Again, usually we want to avoid the conversion, therefore specifying stringsAsFactors=FALSE:
4.4.2 The working directory
When reading files into R, there is another important concept we need to be aware of: the working directory. The R environment always points to a certain directory on our computer, which is knows as the working directory. We can get the current working directory with getwd:
We can set a new working directory with setwd:
When reading a file from the working directory, we can specify just the file name instead of the full path:
When reading and/or writing multiple files from the same directory, it is very convenient to set the working directory at the beginning of our script. That way, in the rest of our script, we can refer to the various files by file name only, rather than by the full path.
4.4.3 Example: the rainfall.csv dataset structure
Let’s read the rainfall.csv file into a data.frame object named rainfall:
This is a longer table than the ones we worked with so far, so printing all of it is inconvenient. Instead, we can use the head or tail function which return a subset of the first or last several rows, respectively:
head(rainfall)
## num altitude sep oct nov dec jan feb mar apr may name
## 1 110050 30 1.2 33 90 117 135 102 61 20 6.7 Kfar Rosh Hanikra
## 2 110351 35 2.3 34 86 121 144 106 62 23 4.5 Saar
## 3 110502 20 2.7 29 89 131 158 109 62 24 3.8 Evron
## 4 111001 10 2.9 32 91 137 152 113 61 21 4.8 Kfar Masrik
## 5 111650 25 1.0 27 78 128 136 108 59 21 4.7 Kfar Hamakabi
## 6 120202 5 1.5 27 80 127 136 95 49 19 2.7 Haifa Port
## x_utm y_utm
## 1 696533.1 3660837
## 2 697119.1 3656748
## 3 696509.3 3652434
## 4 696541.7 3641332
## 5 697875.3 3630156
## 6 687006.2 3633330tail(rainfall)
## num altitude sep oct nov dec jan feb mar apr may name x_utm
## 164 321800 -180 0.2 12 37 55 65 59 36 11 5.0 Sde Eliyahu 736189.3
## 165 321850 -220 0.2 13 33 53 64 56 35 11 4.7 Tirat Zvi 737522.4
## 166 330370 -375 0.1 6 10 20 22 19 11 6 1.3 Kalya 733547.8
## 167 337000 -390 0.0 5 3 10 7 7 7 3 0.5 Sdom 728245.6
## 168 345005 80 0.4 2 2 6 5 4 5 2 0.4 Yotveta 700626.3
## 169 347702 11 0.0 4 2 5 4 3 3 2 1.0 Eilat 689139.3
## y_utm
## 164 3591636
## 165 3590062
## 166 3515345
## 167 3435503
## 168 3307819
## 169 3270290We can also check the table structure with str:
str(rainfall)
## 'data.frame': 169 obs. of 14 variables:
## $ num : int 110050 110351 110502 111001 111650 120202 120630 120750 120870 121051 ...
## $ altitude: int 30 35 20 10 25 5 450 30 210 20 ...
## $ sep : num 1.2 2.3 2.7 2.9 1 1.5 1.9 1.6 1.1 1.8 ...
## $ oct : int 33 34 29 32 27 27 36 31 32 32 ...
## $ nov : int 90 86 89 91 78 80 93 91 93 85 ...
## $ dec : int 117 121 131 137 128 127 161 163 147 147 ...
## $ jan : int 135 144 158 152 136 136 166 170 147 142 ...
## $ feb : int 102 106 109 113 108 95 128 146 109 102 ...
## $ mar : int 61 62 62 61 59 49 71 76 61 56 ...
## $ apr : int 20 23 24 21 21 19 21 22 16 13 ...
## $ may : num 6.7 4.5 3.8 4.8 4.7 2.7 4.9 4.9 4.3 4.5 ...
## $ name : chr "Kfar Rosh Hanikra" "Saar" "Evron" "Kfar Masrik" ...
## $ x_utm : num 696533 697119 696509 696542 697875 ...
## $ y_utm : num 3660837 3656748 3652434 3641332 3630156 ...Create a plot of rainfall in January (
jan) as function of elevation (altitude) based on therainfalltable (Figure 4.4).
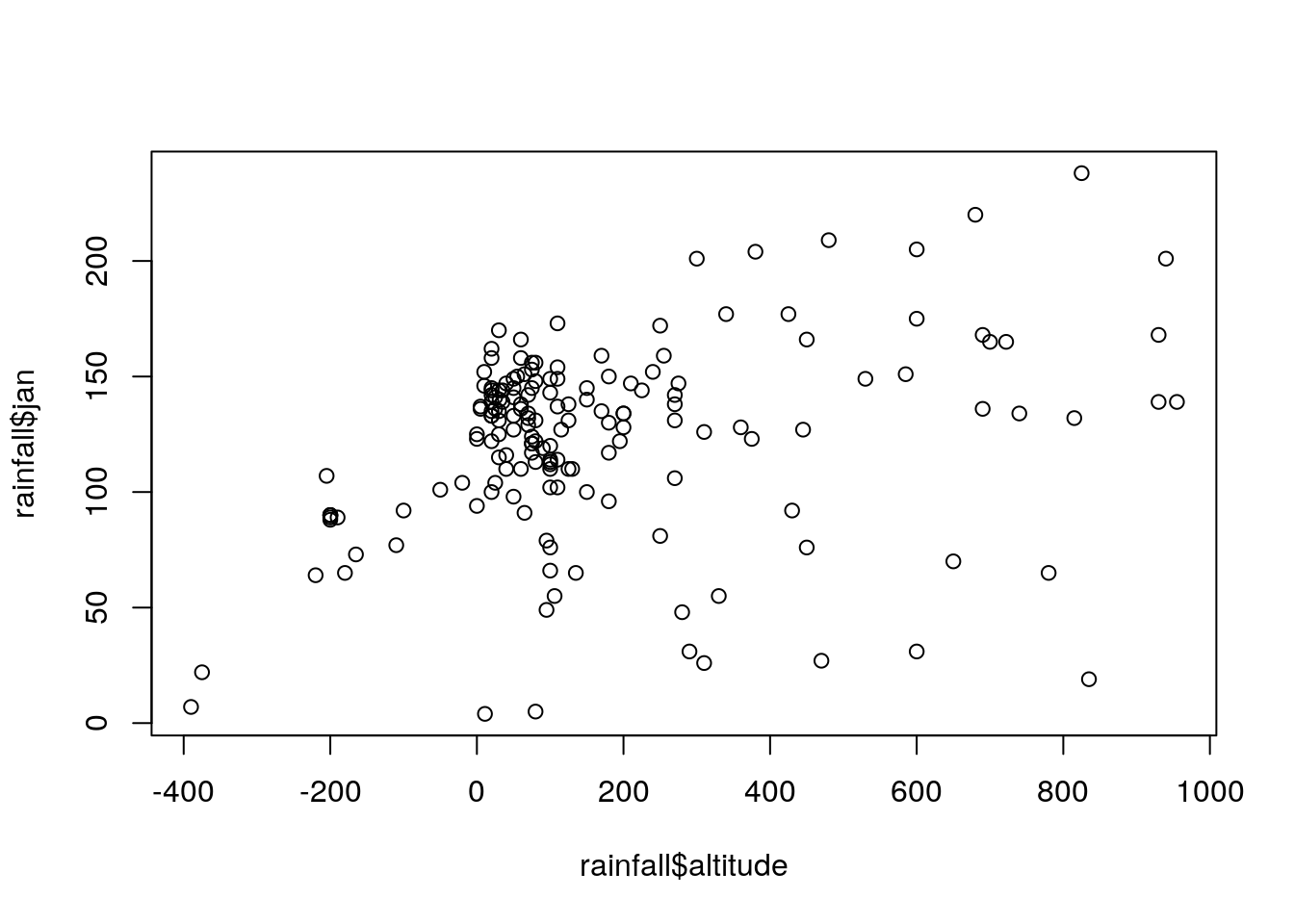
Figure 4.4: Rainfall amount in January as function of elevation
We can get specific information from the table trough subsetting and summarizing. For example, what is the elevation of the lowest and highest stations?
What is the name of the lowest and highest station?
How much rainfall does the "Haifa University" station receive in April?
We can create a new column using assignment (Section 4.1.6). For example, here is how we can create a new column named sep_oct, with the amounts of rainfall in September and October combined:
To accomodate more complex calculations, we can also create a new column inside a for loop, going over table rows. For example, the following code section calculates a new column named annual with the total annual precipitation amounts per station:
m = c("sep", "oct", "nov", "dec", "jan", "feb", "mar", "apr", "may")
for(i in 1:nrow(rainfall)) {
rainfall$annual[i] = sum(rainfall[i, m])
}Go over the above code and make sure you understand how it works.
The updated rainfall table with the new columns is shown below:
head(rainfall)
## num altitude sep oct nov dec jan feb mar apr may name
## 1 110050 30 1.2 33 90 117 135 102 61 20 6.7 Kfar Rosh Hanikra
## 2 110351 35 2.3 34 86 121 144 106 62 23 4.5 Saar
## 3 110502 20 2.7 29 89 131 158 109 62 24 3.8 Evron
## 4 111001 10 2.9 32 91 137 152 113 61 21 4.8 Kfar Masrik
## 5 111650 25 1.0 27 78 128 136 108 59 21 4.7 Kfar Hamakabi
## 6 120202 5 1.5 27 80 127 136 95 49 19 2.7 Haifa Port
## x_utm y_utm sep_oct annual
## 1 696533.1 3660837 34.2 565.9
## 2 697119.1 3656748 36.3 582.8
## 3 696509.3 3652434 31.7 608.5
## 4 696541.7 3641332 34.9 614.7
## 5 697875.3 3630156 28.0 562.7
## 6 687006.2 3633330 28.5 537.24.5 The apply function
In the last example (Section 4.4.3), we basically used a for loop to apply a function (sum) on all rows of a table (rainfall[, m]). The apply function can replace for loops in such situations, and more generally: in situations when we are interested in applying the same function on all subsets of certain dimension of a data.frame, a matrix (Section 5.1.8) or an array (Section 5.2.3).
In case of a data.frame, there are two dimensions that we can work on with apply:
- Rows = Dimension
1 - Columns = Dimension
2
Given the dimension of choice and a function, the apply function splits the table into separate rows or columns, applies the function and combines the results back into a complete object (Figure 4.5). This technique is therefore also known as split-apply-combine.
Figure 4.5: The apply function, applying the mean function on columns (left) or rows (right)
The apply function needs three arguments:
X—The object we are working on:data.frame,matrixorarray15MARGIN—The dimension we are working onFUN—The function applied on that dimension
For example, the apply function can be used, instead of a for loop, to calculate total annual rainfall per station. To do that, we apply the sum function on the rows dimension:
Or, in short:
As another example, we can calculate average monthly rainfall, among all 169 stations, per month. This time, the mean function is applied on the columns dimension:
The result avg_rain is a named numeric vector. Element names correspond to column names of rainfall[, m]:
avg_rain
## sep oct nov dec jan feb mar
## 1.025444 21.532544 64.852071 105.798817 123.053254 103.130178 58.366864
## apr may
## 16.769231 3.968639We can quickly visualize the values with barplot (Figure 4.6):
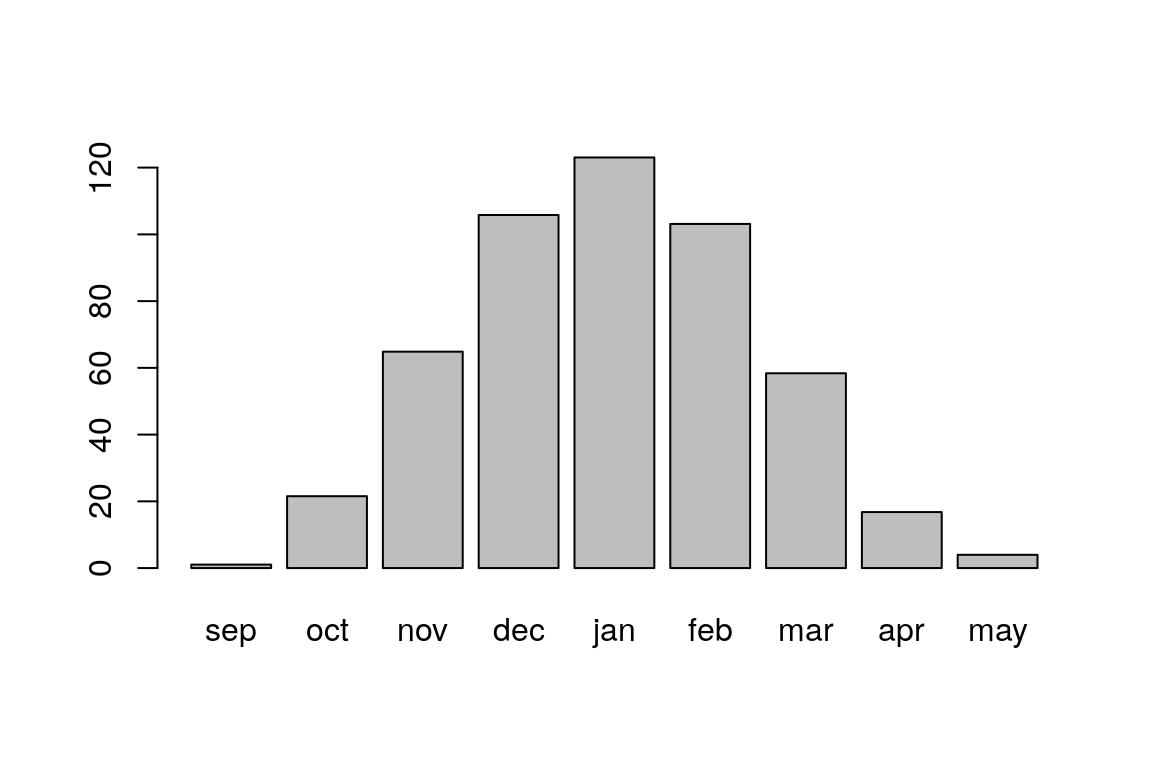
Figure 4.6: Average rainfall per month, among 169 stations in Israel
As another example, let’s use apply to find the station name with the highest rainfall per month. The following expression applies which.max on the columns, returning the row indices where the maximal rainfall values are located per column:
max_st = apply(rainfall[, m], 2, which.max)
max_st
## sep oct nov dec jan feb mar apr may
## 71 23 77 77 77 66 66 66 77We can get the corresponding station names by subsetting the name column using max_st:
rainfall$name[max_st]
## [1] "Eilon" "Maabarot" "Horashim" "Horashim" "Horashim"
## [6] "Golan Farm" "Golan Farm" "Golan Farm" "Horashim"It is convenient to combine the names and values in a table:
4.6 Table joins
4.6.1 Joins for classification
The MOD13A3_2000_2019_dates.csv table contains the dates that each layer in the raster MOD13A3_2000_2019.tif, which we are going to meet in Chapter 5, refers to:
Here is what the first few rows in the dates table look like:
head(dates)
## layer date
## 1 1 2000-02-01
## 2 2 2000-03-01
## 3 3 2000-04-01
## 4 4 2000-05-01
## 5 5 2000-06-01
## 6 6 2000-07-01For further analysis of the raster layers, we would like to be able to group the dates by season. How can we calculate a new season column, specifying the season each date belongs to (Table 4.1)?
| season | months |
|---|---|
"winter" |
12, 1, 2 |
"spring" |
3, 4, 5 |
"summer" |
6, 7, 8 |
"fall" |
9, 10, 11 |
One way to classify dates to seasons is through a combination of subsetting and assignment: assigning each season name into the right subset of a new season column, depending on the month.
First, we need to figure out the month each date belongs to (Section 3.1.2.3):
dates$date = as.Date(dates$date)
dates$month = as.character(dates$date, "%m")
dates$month = as.numeric(dates$month)Now the dates table also contains a month column:
head(dates)
## layer date month
## 1 1 2000-02-01 2
## 2 2 2000-03-01 3
## 3 3 2000-04-01 4
## 4 4 2000-05-01 5
## 5 5 2000-06-01 6
## 6 6 2000-07-01 7Second, we assign the season names according the relevant subset of months:
dates$season[dates$month %in% c(12, 1:2)] = "winter"
dates$season[dates$month %in% 3:5] = "spring"
dates$season[dates$month %in% 6:8] = "summer"
dates$season[dates$month %in% 9:11] = "fall"Here is the result:
head(dates)
## layer date month season
## 1 1 2000-02-01 2 winter
## 2 2 2000-03-01 3 spring
## 3 3 2000-04-01 4 spring
## 4 4 2000-05-01 5 spring
## 5 5 2000-06-01 6 summer
## 6 6 2000-07-01 7 summerThis method of classification may be inconvenient when we have many categories or complex criteria. A more general option is to use a table join (Section 4.6.2).
4.6.2 Joining tables
The merge function can do several types of table joins, including a left join (Figure 4.7). The first two parameters are the tables that need to be joined, x and y. The third by parameter is the common column name(s) by which the tables need to be joined. The parameter all.x=TRUE specifies that all rows of x need to be kept in the resulting table, even if they do not have a match in y, which is the definition of a left join.
![Join types^[http://r4ds.had.co.nz/relational-data.html]](images/lesson_04_join_types.png)
Figure 4.7: Join types16
The table we are going to join with dates is a small table named tab that contains season classification per month. We can prepare the tab table as follows:
tab = data.frame(
month = c(12, 1:11),
season = c(rep("winter", 3), rep("spring", 3), rep("summer", 3), rep("fall", 3)),
stringsAsFactors = FALSE
)
tab
## month season
## 1 12 winter
## 2 1 winter
## 3 2 winter
## 4 3 spring
## 5 4 spring
## 6 5 spring
## 7 6 summer
## 8 7 summer
## 9 8 summer
## 10 9 fall
## 11 10 fall
## 12 11 fallNow we can join the dates and tab tables. Before that, we remove the season column we manually created in the previous example:
dates$season = NULL
head(dates)
## layer date month
## 1 1 2000-02-01 2
## 2 2 2000-03-01 3
## 3 3 2000-04-01 4
## 4 4 2000-05-01 5
## 5 5 2000-06-01 6
## 6 6 2000-07-01 7Then we use merge to join the tables:
Examing the result shows that the season column was indeed joined:
head(dates)
## month layer date season
## 1 1 12 2001-01-01 winter
## 2 1 36 2003-01-01 winter
## 3 1 96 2008-01-01 winter
## 4 1 84 2007-01-01 winter
## 5 1 60 2005-01-01 winter
## 6 1 48 2004-01-01 winterThe joined table was automatically sorted by the common column month. It can be sorted back to chronological order using the order function (Section 2.4.4):
4.7 Writing tables to file
Using write.csv we can write the contents of a data.frame to a CSV file (Figure 4.8):
Figure 4.8: Writing data from the RAM to long-term storage
The row.names parameter determines whether the row names are saved. As mentioned above (Section 4.1.4), data.frame row names are usually meaningless, in which case there is no reason to save them in the CSV file.
Like in read.csv, we can either give a full file path or just the file name. If we specify just the file name, such as in the above example, the file is written to the working directory.